SP-FM (MixFlow)¶
Shortest-Path Flow Matching with Mixture-Conditioned Bases for Out-of-Distribution Generalization to Unseen Conditions.
Home | Installation | Running SP-FM
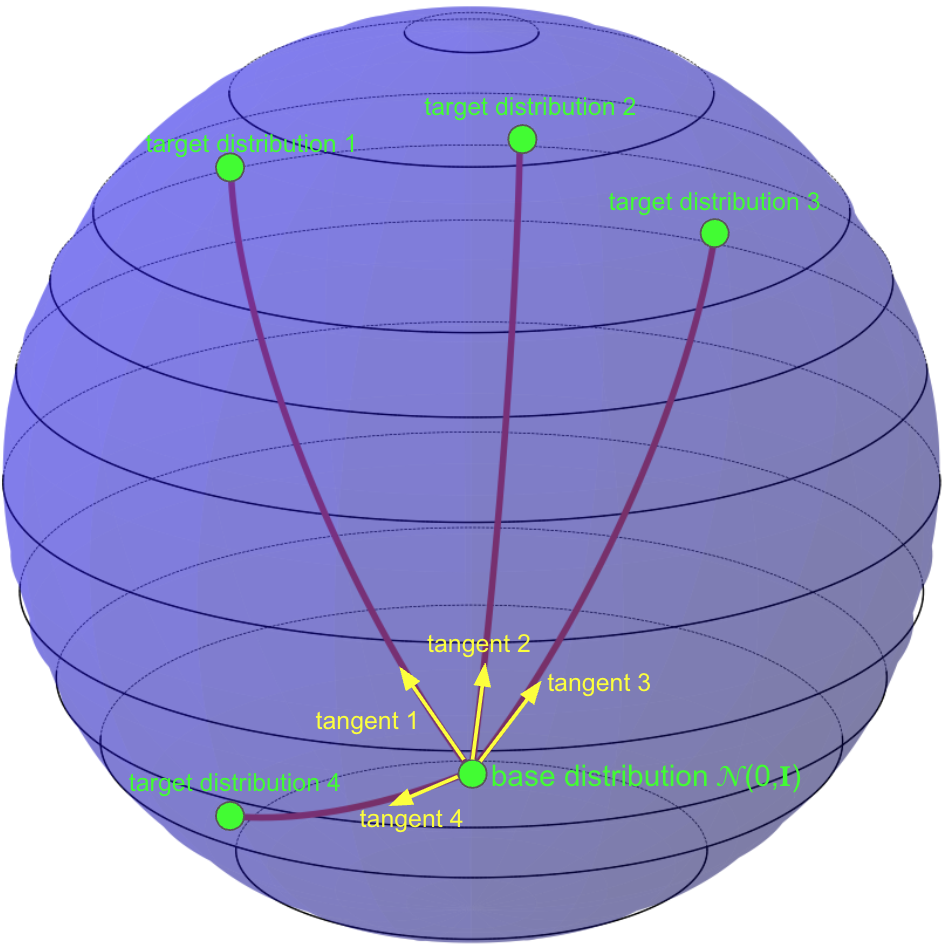
(A) Vanilla CFM |
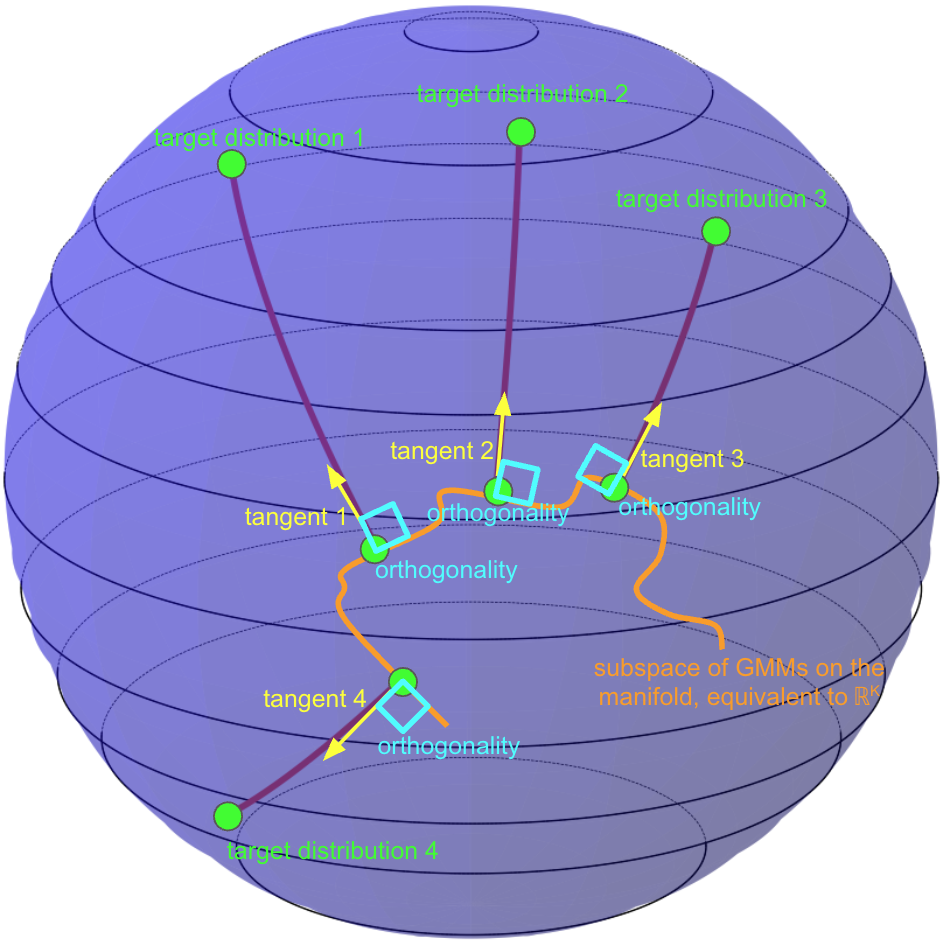
(B) SP-FM |
Overview¶
SP-FM (MixFlow) is a conditional flow-matching framework for descriptor-controlled generation. Instead of relying on a single Gaussian base distribution, SP-FM learns a mixture base and a descriptor-conditioned flow jointly, trained via shortest-path flow matching. This joint modeling is designed to extrapolate smoothly to unseen conditions and improve out-of-distribution generalization across tasks.
Publication¶
This repository accompanies the arXiv manuscript:
- Title: Shortest-Path Flow Matching with Mixture-Conditioned Bases for OOD Generalization to Unseen Conditions
- Authors: Andrea Rubbi, Amir Akbarnejad, Mohammad Vali Sanian, Aryan Yazdan Parast, Hesam Asadollahzadeh, Arian Amani, Naveed Akhtar, Sarah Cooper, Andrew Bassett, Lassi Paavolainen, Pietro Liò, Sattar Vakili, Mo Lotfollahi
- arXiv: 2601.11827v2 [cs.LG] (11 Feb 2026)
Paper link: https://arxiv.org/html/2601.11827v2
Datasets¶
Synthetic Data¶
We construct a synthetic benchmark of letter populations, where each condition corresponds to a letter and a specific rotation. Each descriptor encodes the letter identity and rotation, and SP-FM learns a mixture base distribution per condition. This setup allows us to test extrapolation to unseen letters and rotation angles.
Morphological Perturbations¶
We evaluate SP-FM on high-content imaging data in feature space. Cells (from BBBC021 and RxRx1) are embedded with a vision backbone, and the model is trained to generate unseen phenotypic responses from compound descriptors alone.
Perturbation Datasets¶
For transcriptomic perturbations, we use Chemical- or CRISPR-based single-cell datasets (Norman, Combosciplex, Replogle and iAstrocytes). Conditions correspond to perturbations' embeddings from pretrained models, and SP-FM is trained to model the distribution of perturbed cells.
Gene embeddings from GPT-4 were sourced from GenePert/GenePT data deposit.
URL: https://zenodo.org/records/10833191